MISTI-Switzerland: Computational Modeling Leads to Insights on a Blinding Disease
-
-
slice.mit.edu
- 2
Filed Under
Recommended

Thanks to an MIT International Science and Technology Initiatives (MISTI) internship in summer 2014 and again through MISTI 2.0 during IAP of that year, Zekavat spent a total of four months in Basel working at the Roche Innovation Center Disease Modeling Department and has continued working on the project since then.
The result? Her work on creating a new computational model to understand Age-related Macular Degeneration (AMD) has led to two academic achievements: a podium presentation at the Association for Research in Vision and Opthalmology (ARVO) conference and an accepted first-author manuscript in the Journal of Lipid Research [See the preprint version].
“Our computer model offers new ideas about dry AMD and may help researchers develop new treatments in the future,” she says. “Age-related Macular Degeneration is the leading cause of blindness in elderly people and begins with the un-treatable dry form of AMD in which cholesterol deposits (known as drusen) form behind the retina. To better understand how cholesterol is used in the retina and the reasons that drusen form, we developed a computational model to describe the dynamics of cholesterol in the retina.
"Using mathematical equations, we calculated the amount of cholesterol that is used by the retina on a daily basis. If the used cholesterol is not eliminated from the eye, it will slowly accumulate behind the retina. Our model predicts that it will take many decades for the accumulated cholesterol to reach the size of drusen. This may explain why AMD is seen in the elderly.”
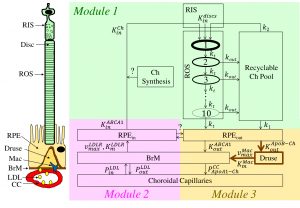 This figure illustrates findings on retinal cholesterol dynamics submitted by Zekavat and her colleagues to the Journal of Lipid Research
This figure illustrates findings on retinal cholesterol dynamics submitted by Zekavat and her colleagues to the Journal of Lipid Research
A key to treatment may come through the next stage of research, she says. “We estimated how fast a drusen deposit can be removed by inflammatory cells called macrophages, which eat up the drusen and eliminate it as small particles that pass into the bloodstream. We found that with enough macrophages, a large drusen can be eliminated in about 10 months, potentially explaining naturally-cleared drusen deposits seen in some individuals. Understanding the processes involved with drusen formation and clearance will help inform treatments for this currently blinding disease.”
Zekavat values her MISTI experiences, both in and out of the lab. She traveled to nearby countries, made new friends, and joined a three-day, 270km bicycle trip around Lake Constantine with the MIT Club of Switzerland.
She continued collaborating on the AMD research after graduation, working with her Roche mentor, clinical pharmacologist and computational modeler Dr. Norman Mazer ’74, MD-PhD ’78, and her MIT MISTI mentor, Thomas Heldt PhD ’04, Helmholtz Career Development Professor of Electrical Engineering & Computer Science.
“The results the RCD model are surprisingly beautiful and also provide insight into possible therapeutic targets for dry AMD. This project showed me how powerful it can be to step back from the wealth of biological literature and ‘connect the dots’ using mathematical modeling. It was also a testament to the power of diversity in research—not only with respect to combining interdisciplinary fields such as math and biology, but also having international and non-academic collaborations in research.”
Inspired by this project, Zekavat has been serving as a computational biologist at the Broad Institute since graduation, continuing her work in the realm of cholesterol metabolism but towards understanding cardiovascular disease through big-data genomics. In the fall she plans to begin pursuing an MD and a PhD in computational biology and bioinformatics in the Yale School of Medicine’s joint degree Medical Scientist Training Program.
Learn more about MISTI, which last year sent 990 students to 25 countries around the world to engage in internships, research, and teaching experiences.








Comments
Emil M Friedman
Fri, 06/02/2017 1:58pm
What effect, if any, do common drugs used to treat blood cholesterol have on the development of dry AMD?
Norman Mazer
Wed, 09/25/2019 11:52am
Model of Retinal Cholesterol Dynamics
Dear Emil,
Just saw your question from 2017. Apologies for the delayed response. As discussed in Maryam's and my publication in the Journal of Lipid Research 2017;58:1325-1337, statin drugs have had no consistent effects on preventing or ameliorating dry AMD. Our paper suggests why. Nonetheless, there is interest in carrying out a large study to see if statins may help some patients with AMD. See Roizenblatt M. et al. Int J Mol Sci. 2018 Nov; 19(11): 3688.
Best regards,
Norm Mazer
'74 SB Physics, '78 MD, Ph.D. Physics